 Herein we describe Neuron Image Analyzer (NIA), an algorithm designed to harness relational information between pixels and thus significantly improve the detection of neuronal structure in images with low SNR. Instead, only pixels with high intensity values determine neuronal structure.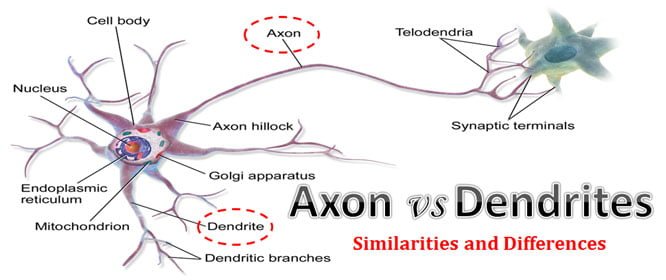  
Current algorithms have difficulty in analyzing the low SNR images because these algorithms do not take into account the relational information of the pixel data. Examples of major issues encountered during image processing of neuronal structures are shown in Fig. The challenge to analyzing neuron images is mainly due to the low signal-to-noise ratio (SNR) of neuronal structures in images obtained from optical microscopy (e.g., bright field, fluorescent microscopy). More importantly, we will show that using these open-source methods can result in the (1) loss of signal associated with neurites, (2) generation of artifact signals and (3) false identification of neuronal structures, all of which prevents precise quantification of changes in neuron morphology post-stimulation. However, methods that use these algorithms 12, 13, 14, 15 still require the merging of two separate sets of images to reveal both immnunostained nuclei and neurites in a neural image and multi-step adjustments to different images (e.g., threshold levels) prior to analysis. Popular algorithms used to analyze neuronal morphology include skeletonization and edge detection 11. These open-source methods use Image J, MATLAB, or Java and have many advantages over commercial software including a reduction in the number of semi-manual annotations required and lower cost. To avoid these issues, many image-processing algorithms are being developed that semi-automatically or automatically detect and quantify the morphology of neurons 10, 11. In addition, the length and direction of neurite extension post-stimulation often is determined by manual tracing, a labor-intensive method that can lead to inconsistent results in repeated measurements. Analyzing images of neurons can be very challenging, however, because neurites are thin (<3.5 μm) arm-like structures and because a high background signal often impedes their accurate visualization. Specifically, the length and direction of neurite extension have been used to quantify the effect of a specific cue on neuronal differentiation, neurite outgrowth and nerve guidance 7, 8, 9. 
We demonstrate that NIA enables precise quantification of neuronal processes (e.g., length and orientation of neurites) in low quality images with a significant increase in the accuracy of detecting neuronal changes post-stimulation.Īnalyzing morphological changes of a nerve cell (i.e., neuron) is one of the key methods for understanding the behavior of neurons in response to various stimuli (e.g., biochemical, electrical, mechanical and topographical) 1, 2, 3, 4, 5, 6. As such, NIA that is based on vector representation is less likely to detect false signals (i.e., non-neuronal structures) or generate artifact signals (i.e., deformation of original structures) than current image analysis algorithms that are based on raster representation. 
In this paper, we describe Neuron Image Analyzer (NIA), a novel algorithm that overcomes these inadequacies by employing Laplacian of Gaussian filter and graphical models (i.e., Hidden Markov Model, Fully Connected Chain Model) to specifically extract relational pixel information corresponding to neuronal structures (i.e., soma, neurite). This inadequacy derives from the fact that these methods often include data from non-neuronal structures or artifacts by simply tracing pixels with high intensity. Both manual and automated methods in current use are severely inadequate at detecting and quantifying changes in neuronal morphology when the images analyzed have a low signal-to-noise ratio (SNR). Image analysis software is an essential tool used in neuroscience and neural engineering to evaluate changes in neuronal structure following extracellular stimuli. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
